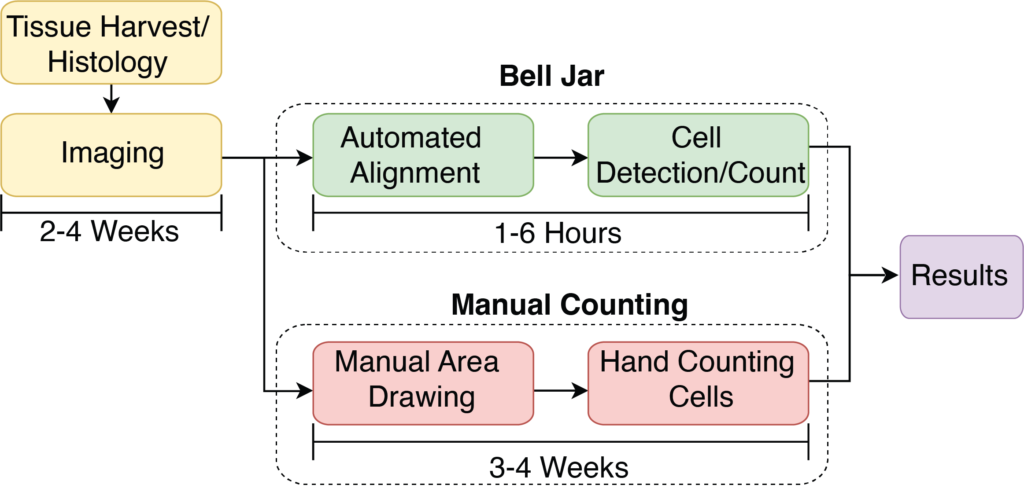
Bell Jar: A Semi-Automated Registration and Cell Counting Tool for Mouse Neurohistology Analysis
Soronow A, Jacobs MW, Dickson RG, Kim EJ. (2022)

Fatherhood is life changing: Uncovering structural and functional changes in the dad brain
Dickson RG, Jacobs MW, Kim EJ. (2022)
Monosynaptic Projections to Excitatory and Inhibitory preBötzinger Complex Neurons
Yang CF, Kim EJ, Callaway EM, Feldman, JL. (2020)
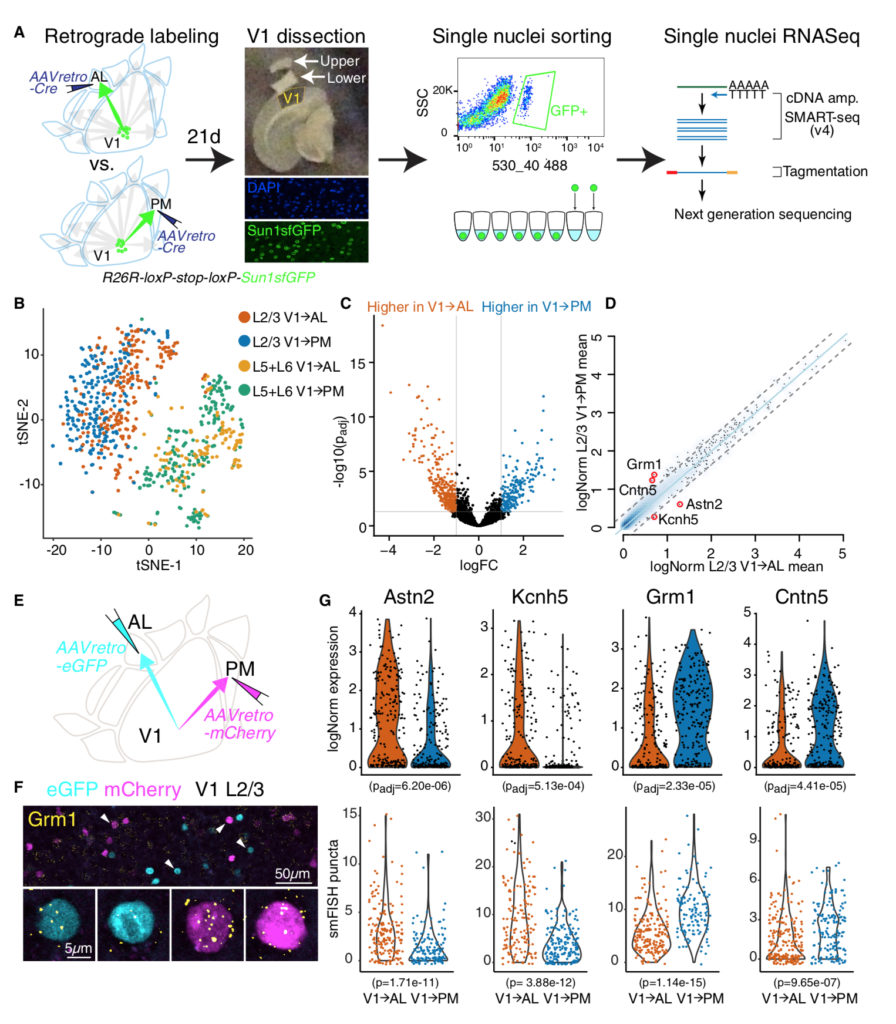
Extraction of distinct neuronal cell types from within a genetically continuous population
Kim EJ, Zhang Z, Huang L, Ito-Cole T, Jacobs MW, Juavinett AL, Senturk G, Hu M, Ku M, Ecker JR, Callaway EM. (2020)
A systematic topographical relationship between mouse lateral posterior thalamic neurons and their visual cortical projection targets
Juavinett AL, Kim EJ, Collins HC, Callaway EM. (2020)
In vivo genome editing via CRISP/Cas9 mediated homology-independent targeted integration
Suzuki K, Tsunekawa Y, Hernandez-Benitez R, Wu J, Zhu J, Kim EJ, Hatanaka F, Yamamoto M, Araoka T, Li Z, Kurita M, Hishida T, Li M, Aizawa E, Guo S, Chen S, Goebl A, Soligalla R, Qu J, Jiang T, Fu X, Jafari M, Esteban CR, Lajara J, Nuñez E, Guillen P, Campistol JM, Matsuzaki F, Liu, G Magistretti P, Zhang K, Callaway EM, Zang K, Izpisua Belmonte JC. (2016)
Improved Monosynaptic Neural Circuit Tracing Using Engineered Rabies Virus Glycoproteins
Kim EJ, Jacobs MW, Ito-Cole T, Callaway EM. (2016)
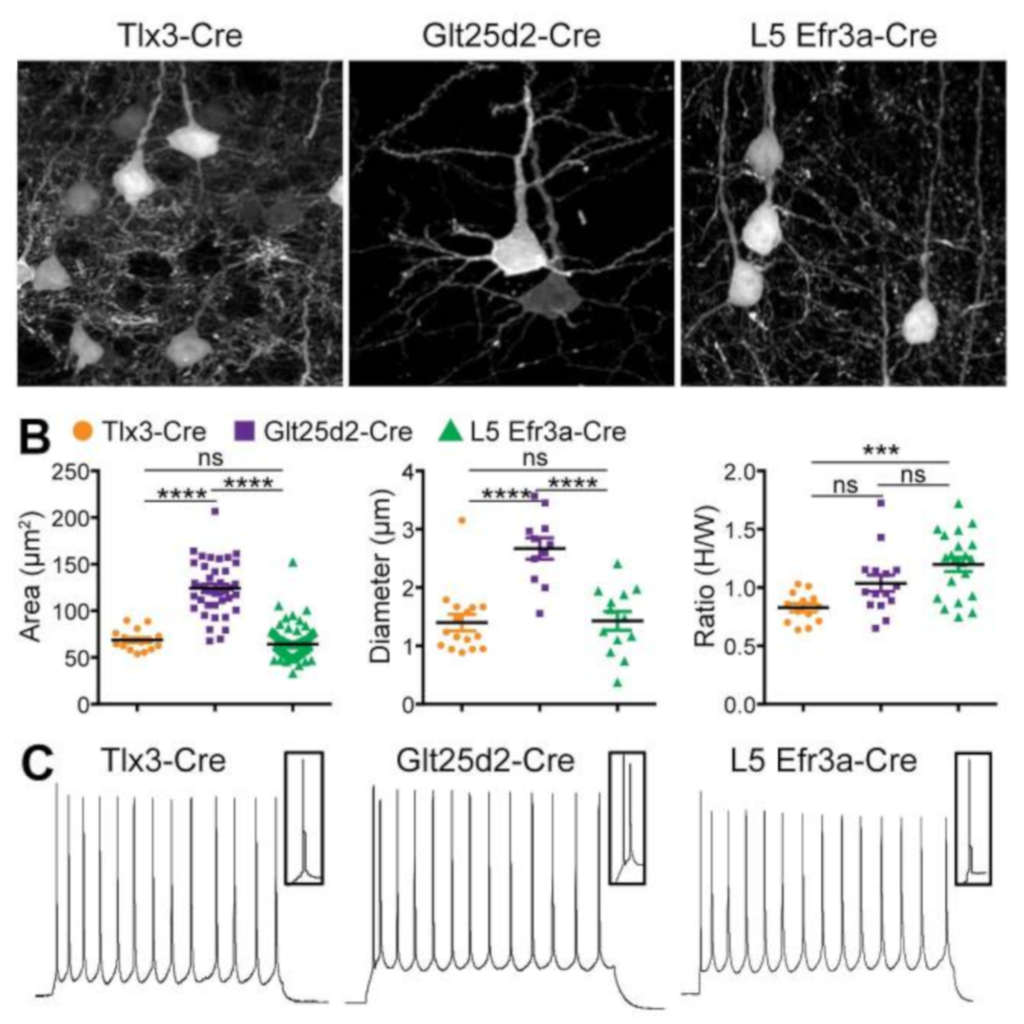
Three Types of Cortical Layer 5 Neurons That Differ in Brain-wide Connectivity and Function
Kim EJ, Juavinett AL, Kyubwa EM, Jacobs MW, Callaway EM. (2015)
Gate control of mechanical itch by a subpopulation of spinal cord interneurons
Bourane S, Duan B, Koch SC, Dalet A, Britz O, Garcia-Campmany L, Kim E, Cheng L, Ghosh A, Ma Q, Goulding M. (2015)
Adult lineage-restricted CNS progenitors specify distinct glioblastoma subtypes
Alcantara Llaguno SR, Wang Z, Sun D, Chen J, Xu J, Kim E, Hatanpaa KJ, Raisanen JM, Burns DK, Johnson JE, Parada LF (2015)
Ascl1 controls the number and distribution of astrocytes and oligodendrocytes in the gray matter and white matter of the spinal cord
Vue TY, Kim EJ, Parras CM, Guillemot F, Johnson JE. (2014)
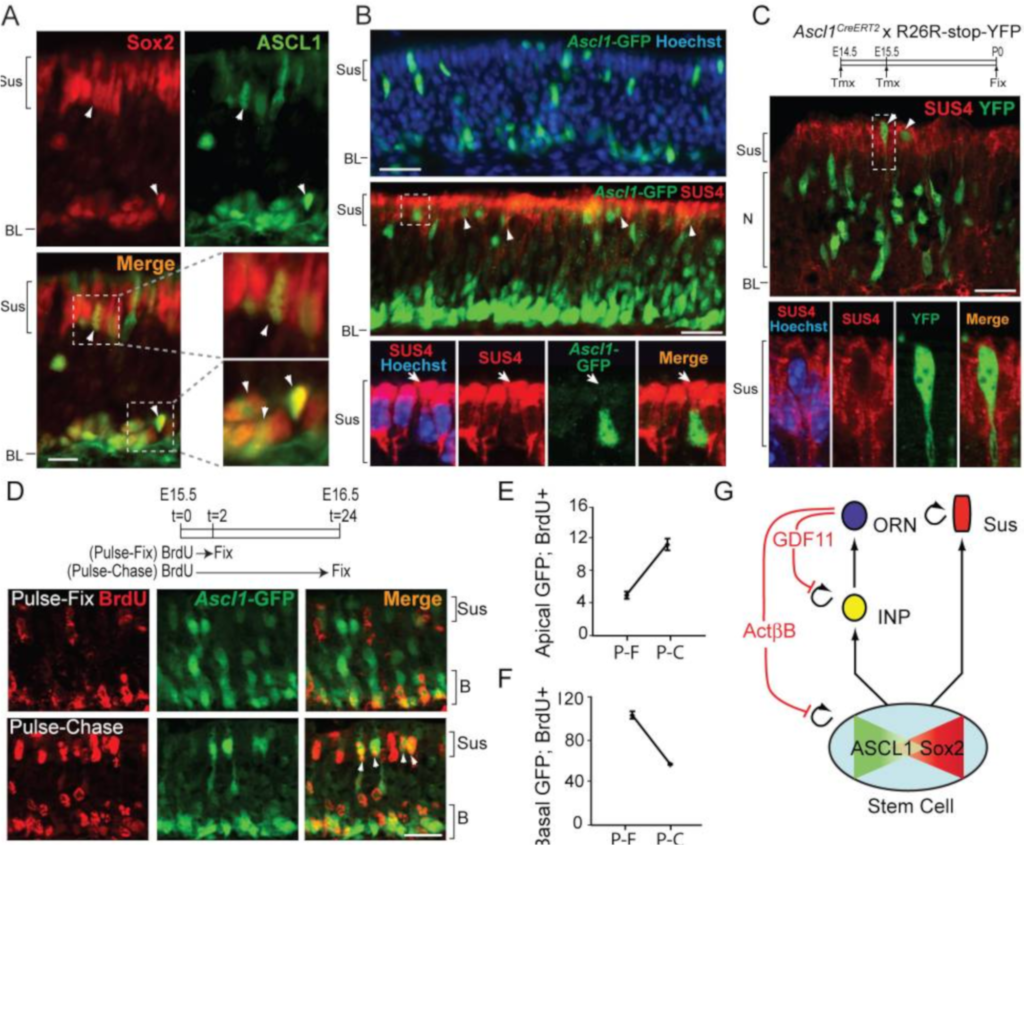
Feedback by Activin and Gdf11 couples regulation of proliferative control and fate choice of neuroepithelial stem/progenitor cells
Gokoffski KK, Wu H, Beites CL, Kim J, Kim EJ, Matzuk MM, Johnson JE, Lander AD, Calof AL (2011)
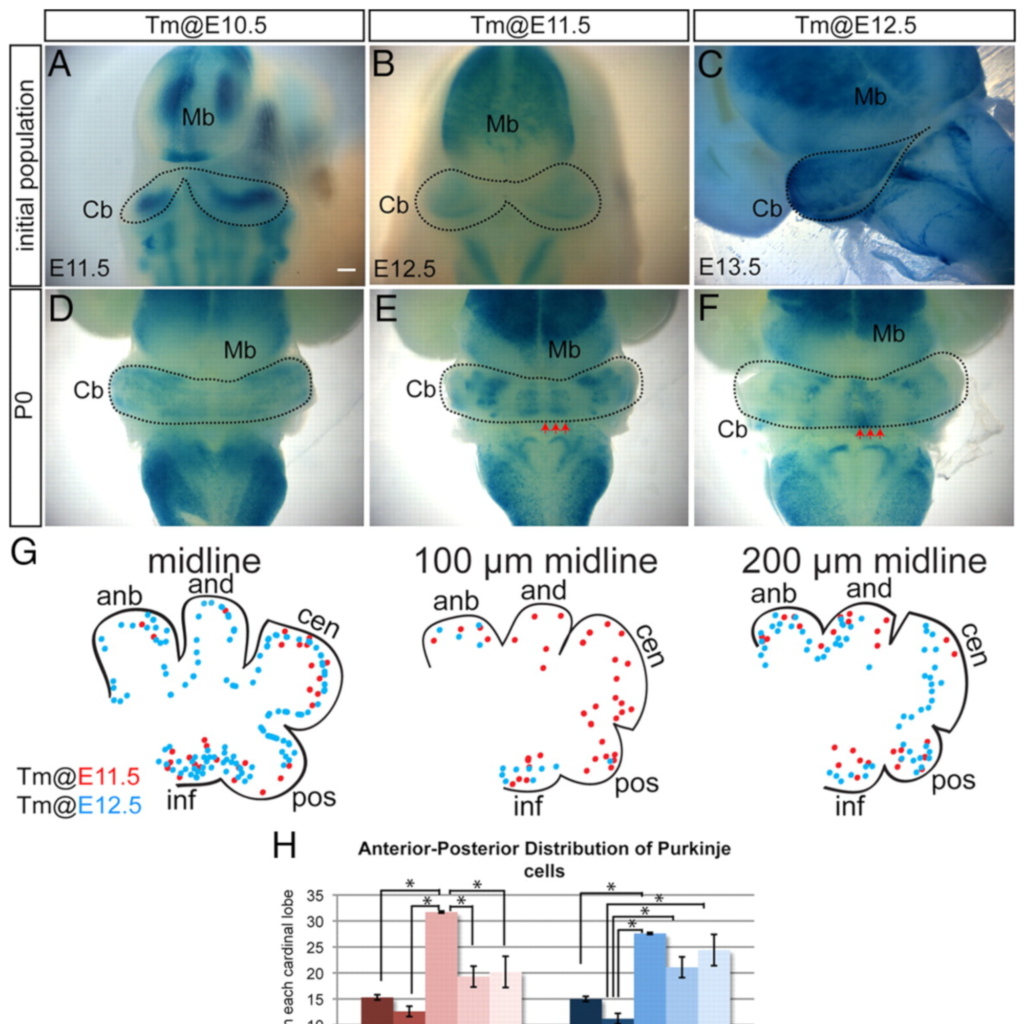
Ascl1 genetics reveals insights into cerebellum local circuit assembly
Sudarov A, Turnbull R, Kim EJ, Guillemot F, Joyner AL (2011)
Ascl1 expression defines a subpopulation of lineage-restricted progenitors in the mammalian retina
Brzezinki JA 4th, Kim EJ, Johnson JE, Reh TA (2011)
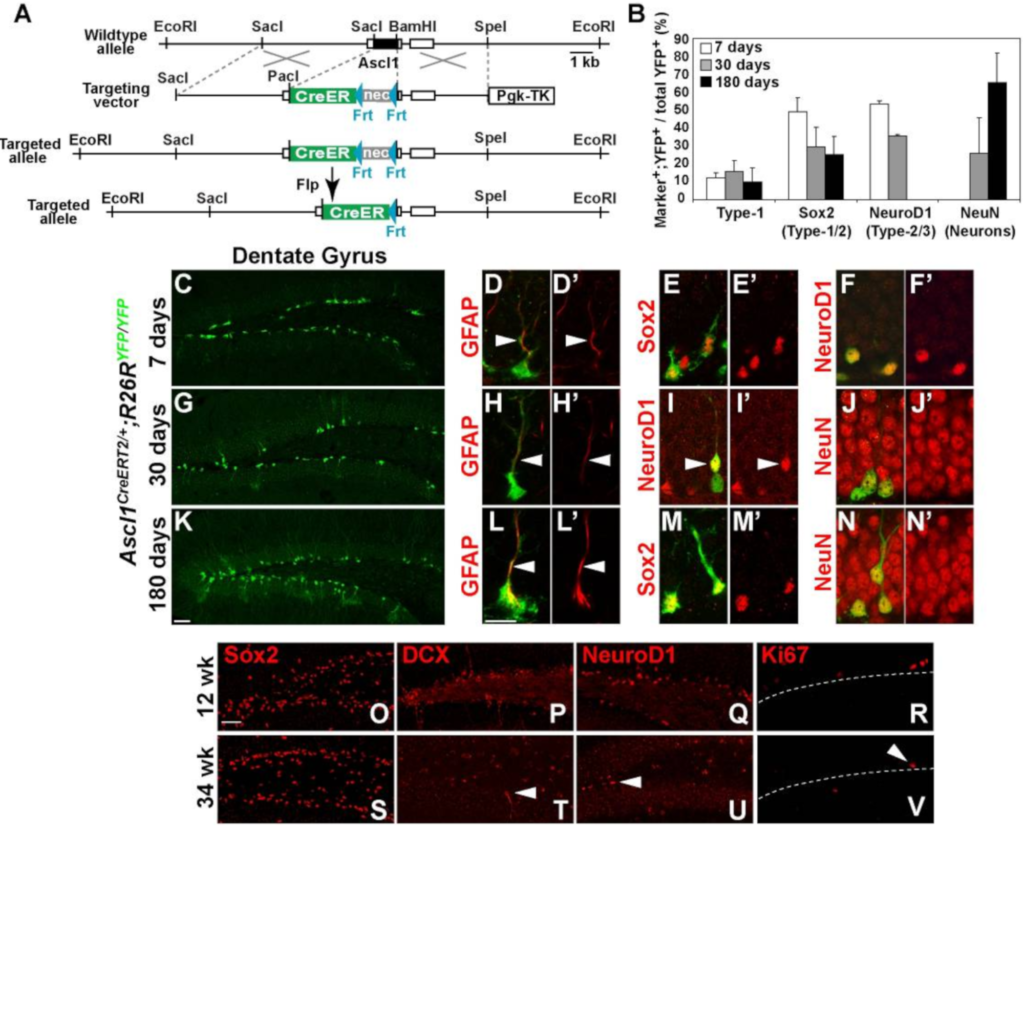
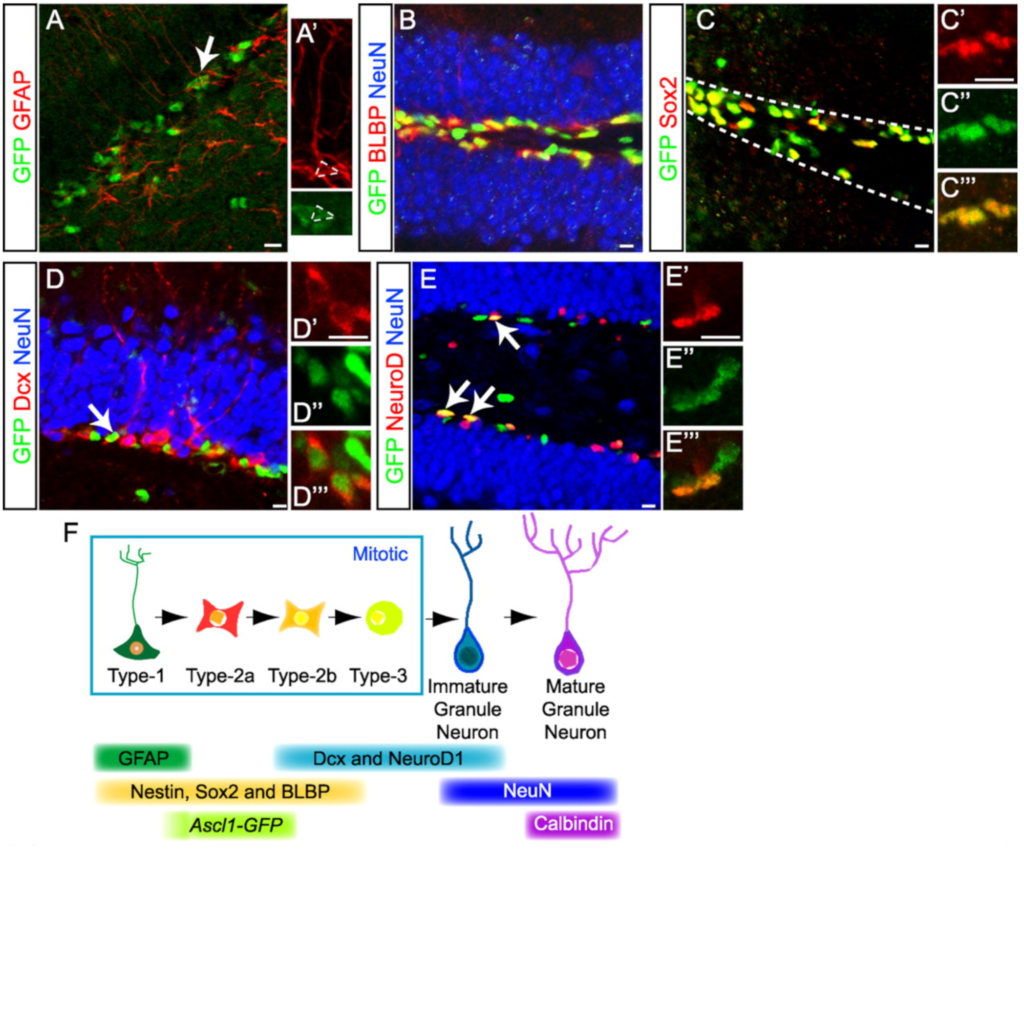
Ascl1 (Mash1) defines cells with long-term neurogenic potential in adult mouse subgranular and subventricular zones
Kim EJ, Ables JA, Dickel LK, Eisch AJ, Johnson JE (2011)
The spatiotemporal fate map of neurogenin1 (Neurog1) lineages in the mouse central nervous system
Kim EJ, Hori K, Wyckoff A, Dickel LK, Koundakjian EJ, Goodrich LV, Johnson JE (2011)
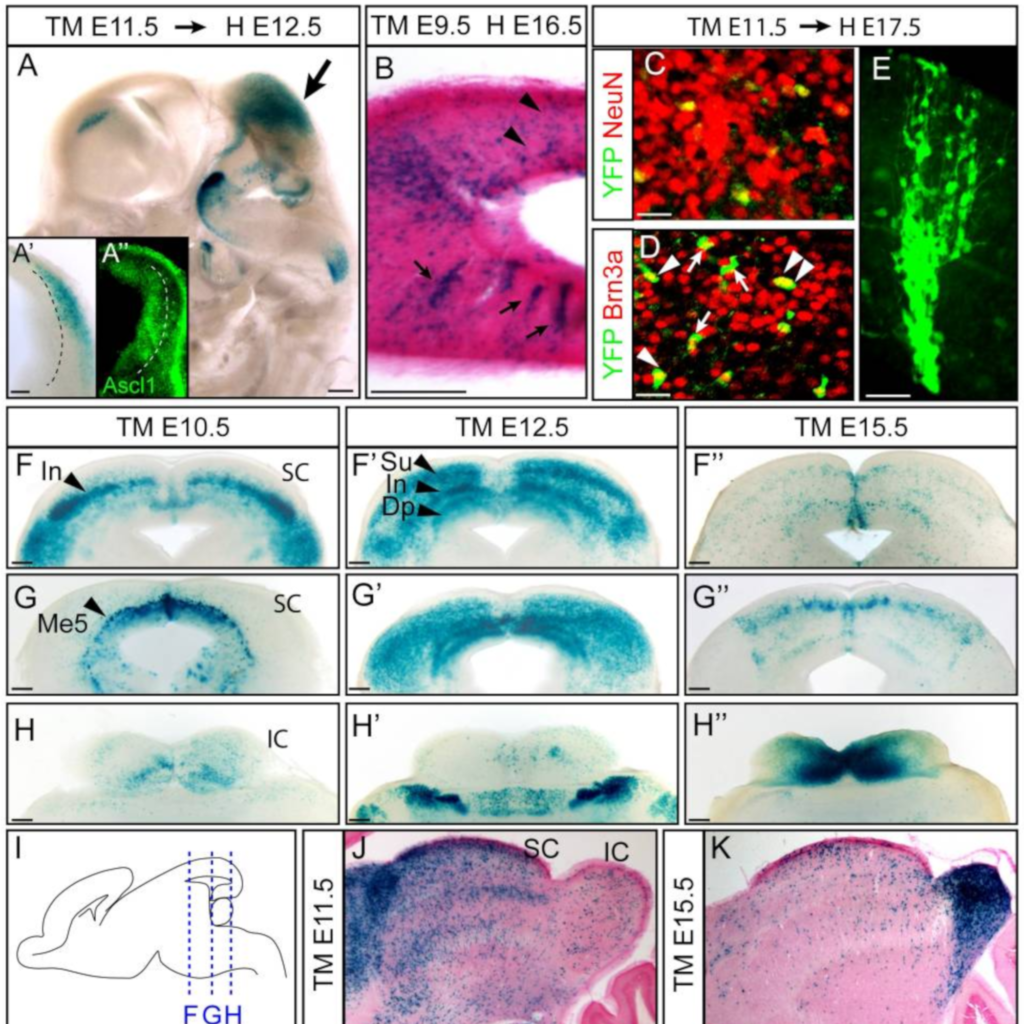
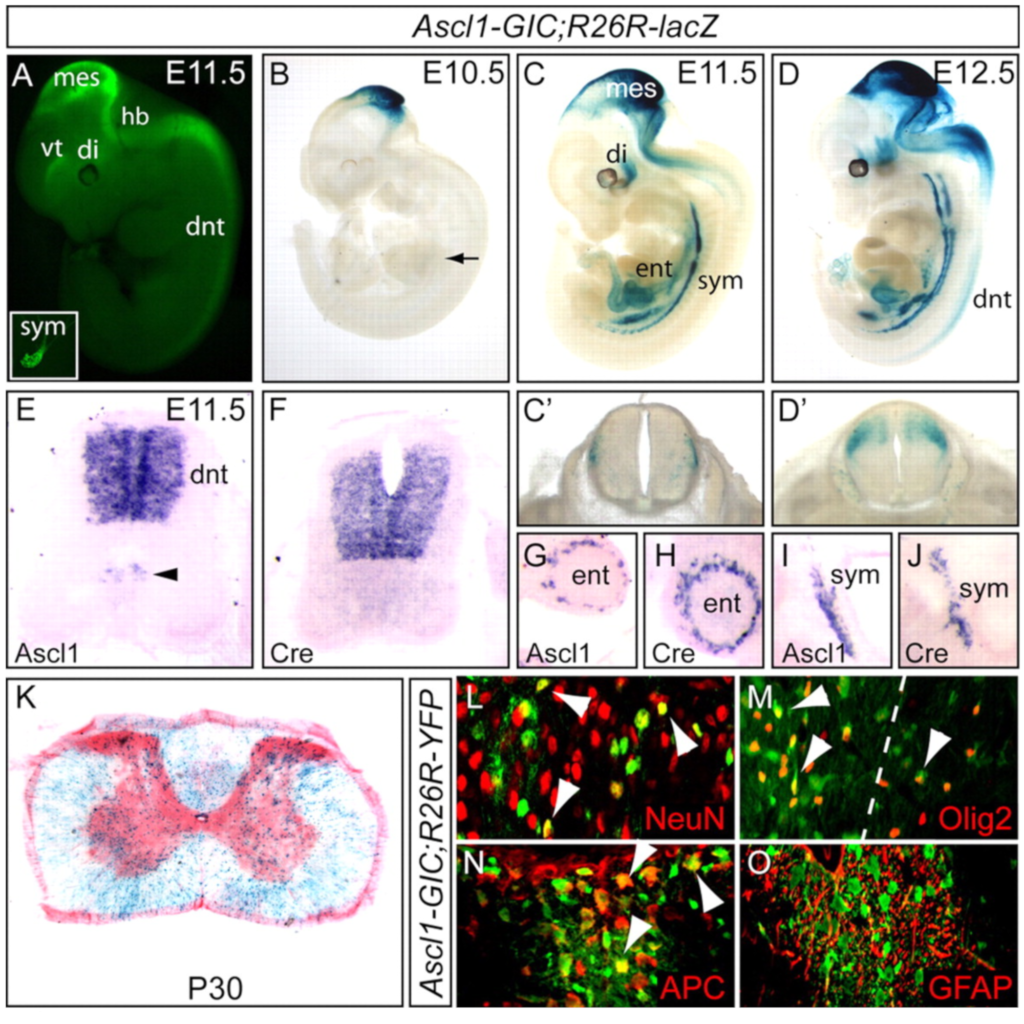
Ascl1 (Mash1) lineage cells contribute to discrete cell populations in CNS
Kim EJ, Battiste J, Nakagawa Y, Johnson JE (2008)
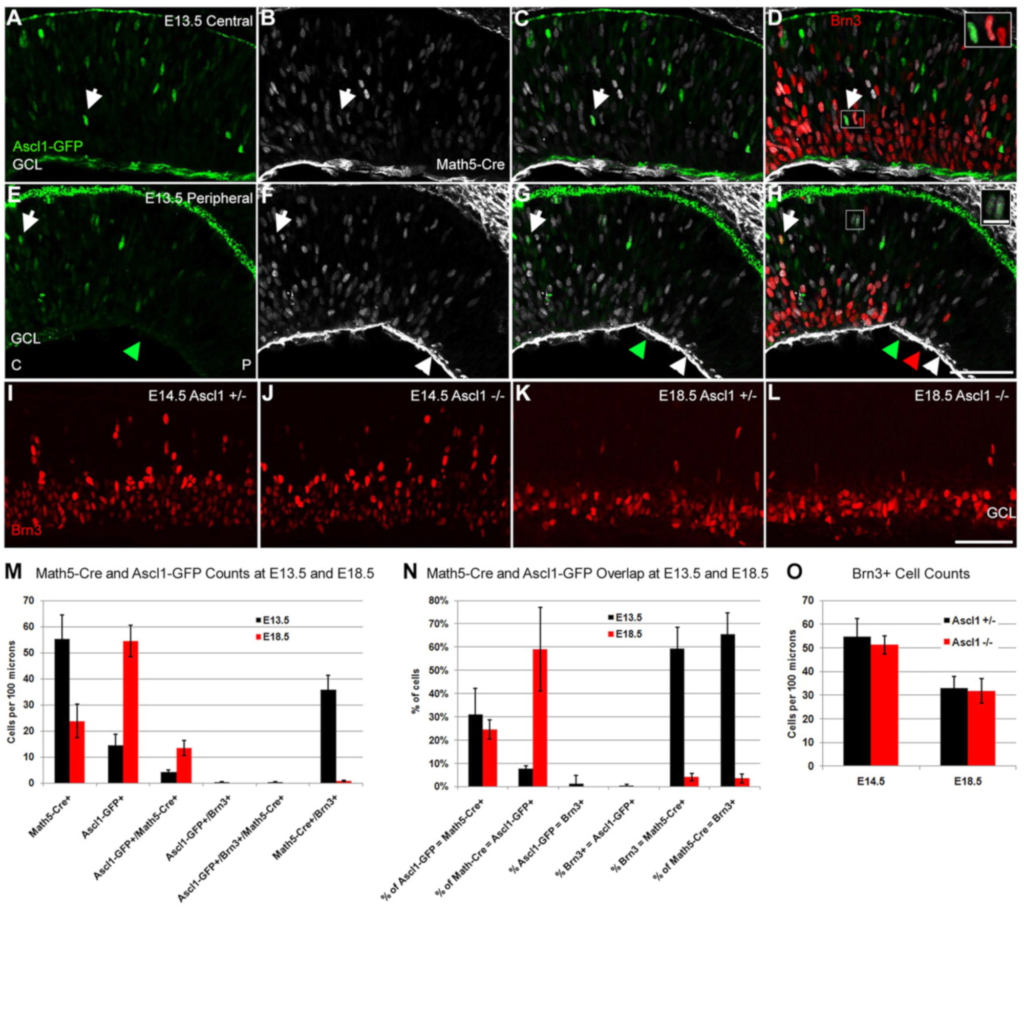
In vivo analysis of Ascl1 defined progenitors reveals distinct developmental dynamics during adult neurogenesis and gliogenesis
Kim EJ, Leung CT, Reed RR, Johnson JE (2007)
Ascl1 defines sequentially generated lineage-restricted neuronal and oligodendrocyte precursor cells in the spinal cord
Battiste J, Helms AW, Kim EJ, Savage TK, Lagace DC, Mandyam CD, Eisch AJ, Miyoshi G, Johnson JE (2007)